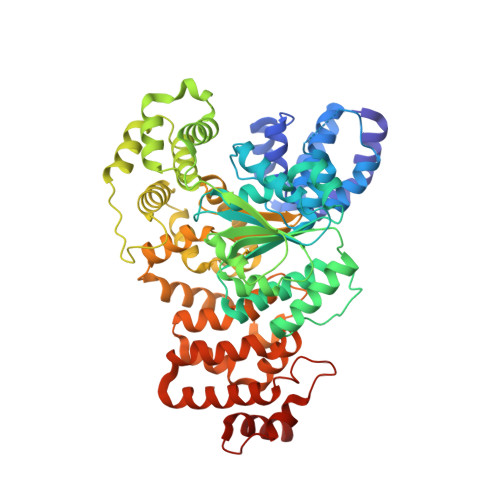
Structure of the apoptotic protease-activating factor 1 bound to ADP
Riedl, S.J., Li, W., Chao, Y., Schwarzenbacher, R., Shi, Y.(2005) Nature 434: 926-933
- PubMed: 15829969
- DOI: https://doi.org/10.1038/nature03465
- Primary Citation of Related Structures:
1Z6T - PubMed Abstract:
Apoptosis is executed by caspases, which undergo proteolytic activation in response to cell death stimuli. The apoptotic protease-activating factor 1 (Apaf-1) controls caspase activation downstream of mitochondria. During apoptosis, Apaf-1 binds to cytochrome c and in the presence of ATP/dATP forms an apoptosome, leading to the recruitment and activation of the initiator caspase, caspase-9 (ref. 2). The mechanisms underlying Apaf-1 function are largely unknown. Here we report the 2.2-A crystal structure of an ADP-bound, WD40-deleted Apaf-1, which reveals the molecular mechanism by which Apaf-1 exists in an inactive state before ATP binding. The amino-terminal caspase recruitment domain packs against a three-layered alpha/beta fold, a short helical motif and a winged-helix domain, resulting in the burial of the caspase-9-binding interface. The deeply buried ADP molecule serves as an organizing centre to strengthen interactions between these four adjoining domains, thus locking Apaf-1 in an inactive conformation. Apaf-1 binds to and hydrolyses ATP/dATP and their analogues. The binding and hydrolysis of nucleotides seem to drive conformational changes that are essential for the formation of the apoptosome and the activation of caspase-9.
Organizational Affiliation:
Department of Molecular Biology, Princeton University, Lewis Thomas Laboratory, Washington Road, Princeton, New Jersey 08544, USA.