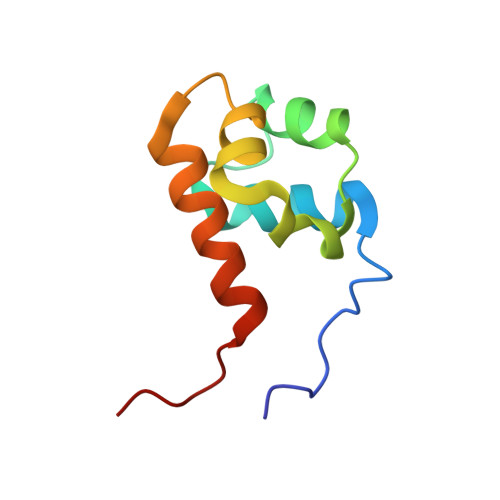
Structure of the Sterile {Alpha} Motif (Sam) Domain of the Saccharomyces Cerevisiae Mitogen-Activated Protein Kinase Pathway-Modulating Protein Ste50 and Analysis of its Interaction with the Ste11 Sam
Grimshaw, S.J., Mott, H.R., Stott, K.M., Nielsen, P.R., Evetts, K.A., Hopkins, L.J., Nietlispach, D., Owen, D.(2004) J Biol Chem 279: 2192
- PubMed: 14573615
- DOI: https://doi.org/10.1074/jbc.M305605200
- Primary Citation of Related Structures:
1UQV - PubMed Abstract:
The sterile alpha motif (SAM) is a 65-70-amino acid domain found in over 300 proteins that are involved in either signal transduction or transcriptional activation and repression. SAM domains have been shown to mediate both homodimerization and heterodimerization and in some cases oligomerization. Here, we present the solution structure of the SAM domain of the Saccharomyces cerevisiae protein, Ste50p. Ste50p functions as a modulator of the mitogen-activated protein kinase (MAPK) cascades in S. cerevisiae, which control mating, pseudohyphal growth, and osmo-tolerance. This is the first example of the structure of a SAM domain from a MAPK module protein. We have studied the associative behavior of Ste50p SAM in solution and shown that it is monomeric. We have examined the SAM domain from Ste11p, the MAPK kinase kinase that associates with Ste50p in vivo, and shown that it forms dimers with a self-association K(d) of approximately 0.5 mm. We have also analyzed the interaction of Ste50p SAM with Ste11p SAM and the effects of mutations at Val-37, Asp-38, Pro-71, Leu-73, Leu-75, and Met-99 of STE50 on the heterodimerization properties of Ste50p SAM. We have found that L73A and L75A abrogate the Ste50p interaction with Ste11p, and we compare these data with the known interaction sites defined for other SAM domain interactions.
Organizational Affiliation:
Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, United Kingdom.