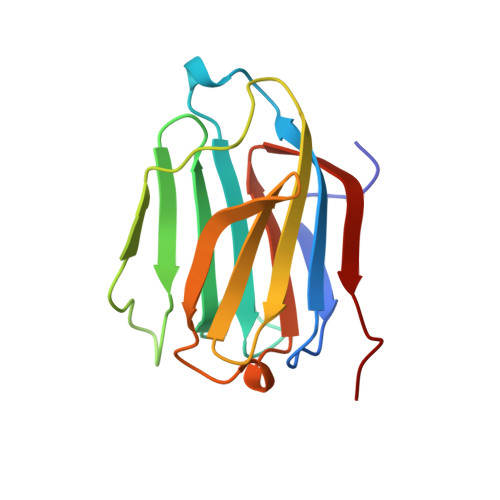
Selective recognition of mannose by the human eosinophil Charcot-Leyden crystal protein (galectin-10): a crystallographic study at 1.8 A resolution.
Swaminathan, G.J., Leonidas, D.D., Savage, M.P., Ackerman, S.J., Acharya, K.R.(1999) Biochemistry 38: 13837-13843
- PubMed: 10529229
- DOI: https://doi.org/10.1021/bi990756e
- Primary Citation of Related Structures:
1QKQ - PubMed Abstract:
The role(s) of the eosinophil Charcot-Leyden crystal (CLC) protein in eosinophil or basophil function or associated inflammatory processes is yet to be established. Although the CLC protein has been reported to exhibit weak lysophospholipase activity, it shows virtually no sequence homology to any known member of this family of enzymes. The X-ray crystal structure of the CLC protein is very similar to the structure of the galectins, members of a beta-galactoside-specific animal lectin family, including a partially conserved galectin carbohydrate recognition domain (CRD). In the absence of any known natural carbohydrate ligand for this protein, the functional role of the CLC protein (galectin-10) has remained speculative. Here we describe structural studies on the carbohydrate binding properties of the CLC protein and report the first structure of a carbohydrate in complex with the protein. Interestingly, the CLC protein demonstrates no affinity for beta-galactosides and binds mannose in a manner very different from those of other related galectins that have been shown to bind lactosamine. The partial conservation of residues involved in carbohydrate binding led to significant changes in the topology and chemical nature of the CRD, and has implications for carbohydrate recognition by the CLC protein in vivo and its functional role in the biology of inflammation.
Organizational Affiliation:
Department of Biology and Biochemistry, University of Bath, Claverton Down, U.K.