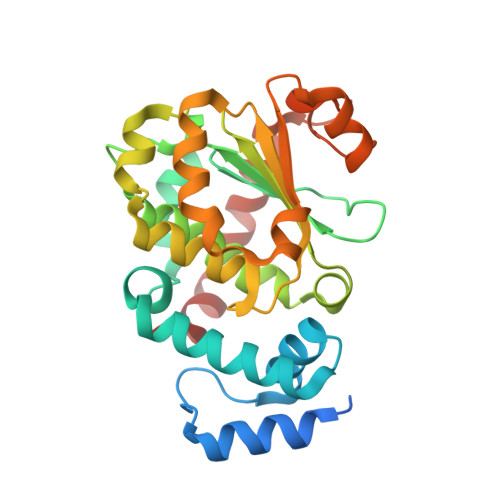
The Molecular Basis of Vitamin E Retention: Structure of Human Alpha-Tocopherol Transfer Protein
Meier, R., Tomizaki, T., Schulze-Briese, C., Baumann, U., Stocker, A.(2003) J Mol Biol 331: 725
- PubMed: 12899840
- DOI: https://doi.org/10.1016/s0022-2836(03)00724-1
- Primary Citation of Related Structures:
1OIP, 1OIZ - PubMed Abstract:
Alpha-tocopherol transfer protein (alpha-TTP) is a liver protein responsible for the selective retention of alpha-tocopherol from dietary vitamin E, which is a mixture of alpha, beta, gamma, and delta-tocopherols and the corresponding tocotrienols. The alpha-TTP-mediated transfer of alpha-tocopherol into nascent VLDL is the major determinant of plasma alpha-tocopherol levels in humans. Mutations in the alpha-TTP gene have been detected in patients suffering from low plasma alpha-tocopherol and ataxia with isolated vitamin E deficiency (AVED). The crystal structure of alpha-TTP reveals two conformations. In its closed tocopherol-charged form, a mobile helical surface segment seals the hydrophobic binding pocket. In the presence of detergents, an open conformation is observed, which probably represents the membrane-bound form. The selectivity of alpha-TTP for RRR-alpha-tocopherol is explained from the van der Waals contacts occurring in the lipid-binding pocket. Mapping the known mutations leading to AVED onto the crystal structure shows that no mutations occur directly in the binding pocket.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Berne, Freiestrasse 3, 3012 Berne, Switzerland.