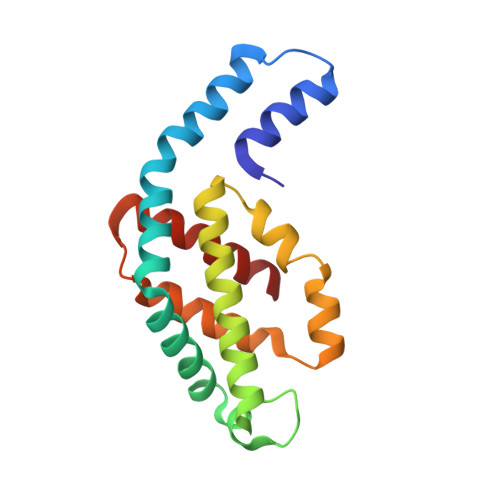
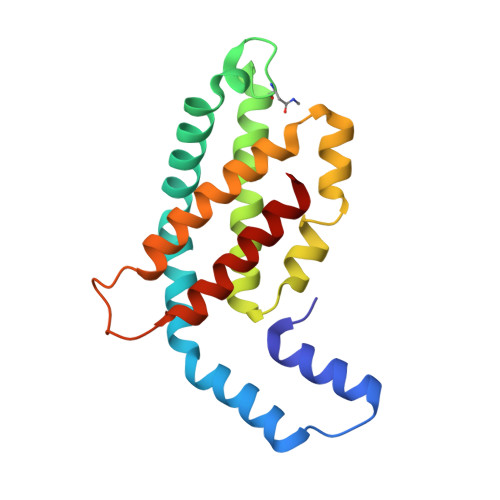
Refined structure of c-phycocyanin from the cyanobacterium Synechococcus vulcanus at 1.6 A: insights into the role of solvent molecules in thermal stability and co-factor structure
Adir, N., Vainer, R., Lerner, N.(2002) Biochim Biophys Acta 1556: 168-174
- PubMed: 12460674
- DOI: https://doi.org/10.1016/s0005-2728(02)00359-6
- Primary Citation of Related Structures:
1KTP - PubMed Abstract:
The crystal structure of the light-harvesting phycobiliprotein, c-phycocyanin from the thermophilic cyanobacterium Synechococcus vulcanus has been refined to 1.6 A resolution based on the previously determined lower resolution structure (PDB entry 1I7Y). The improved data was collected using synchrotron radiation at 100 K. The significantly improved crystallographic data has lead to improved calculated electron density maps, allowing the unambiguous positioning of all protein and co-factor atoms and the positioning of 377 solvent molecules. The positions of solvent molecules at specific sites important for stabilization of different levels of self-assembly of the phycobilisome structure were identified and the bonding network is described. The presence of solvent molecules in the vicinity of the co-factors and in intermolecular spaces is identified and their possible roles are suggested. All three of the phycocyanobilin co-factors bind water molecules at specific sites between the propionic acid side chains. Molecular dynamic (MD) simulations support that these special waters have a role in stabilization of this conformation. On the basis of the crystal packing reported here and in comparison to other phycobiliprotein crystal forms, we have analyzed the roles of specific sites on the formation of the phycobilisome complex.
Organizational Affiliation:
Department of Chemistry and Institute of Catalysis, Science and Technology, Technion-Israel Institute of Technology, City, Haifa 32000, Technion, Israel. nadir@tx.technion.ac.il