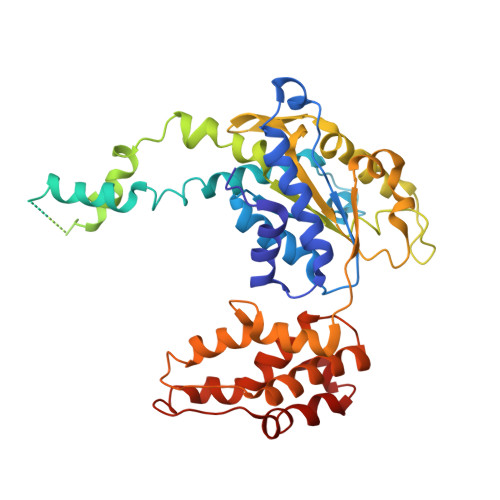
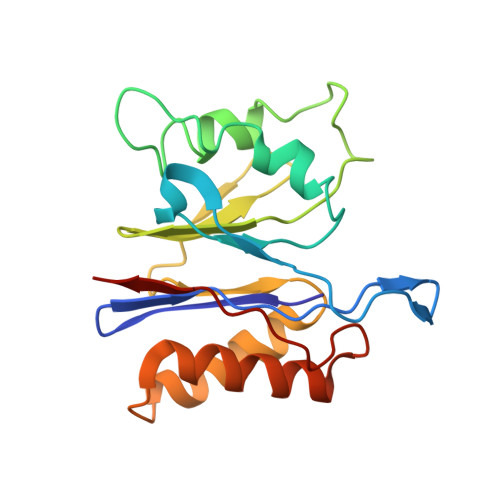
Crystal structures of the HslVU peptidase-ATPase complex reveal an ATP-dependent proteolysis mechanism.
Wang, J., Song, J.J., Franklin, M.C., Kamtekar, S., Im, Y.J., Rho, S.H., Seong, I.S., Lee, C.S., Chung, C.H., Eom, S.H.(2001) Structure 9: 177-184
- PubMed: 11250202
- DOI: https://doi.org/10.1016/s0969-2126(01)00570-6
- Primary Citation of Related Structures:
1G4A, 1G4B - PubMed Abstract:
The bacterial heat shock locus HslU ATPase and HslV peptidase together form an ATP-dependent HslVU protease. Bacterial HslVU is a homolog of the eukaryotic 26S proteasome. Crystallographic studies of HslVU should provide an understanding of ATP-dependent protein unfolding, translocation, and proteolysis by this and other ATP-dependent proteases.
Organizational Affiliation:
Department of Molecular Biophysics, Biochemistry, 266 Whitney Avenue, Yale University, 06520, New Haven, CT, USA. wang@mail.csb.yale.edu