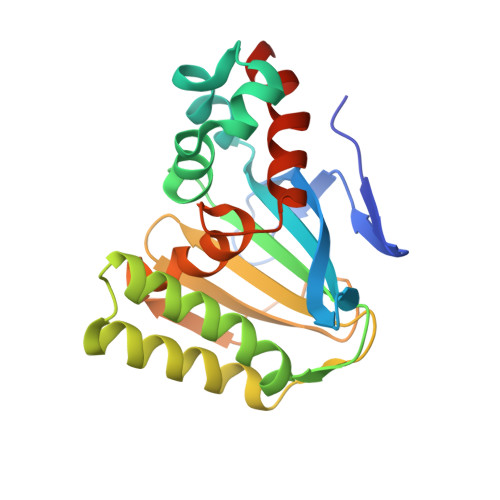
Structure and mechanism of the evolutionarily unique plant enzyme chalcone isomerase.
Jez, J.M., Bowman, M.E., Dixon, R.A., Noel, J.P.(2000) Nat Struct Biol 7: 786-791
- PubMed: 10966651
- DOI: https://doi.org/10.1038/79025
- Primary Citation of Related Structures:
1EYP, 1EYQ - PubMed Abstract:
Chalcone isomerase (CHI) catalyzes the intramolecular cyclization of chalcone synthesized by chalcone synthase (CHS) into (2S)-naringenin, an essential compound in the biosynthesis of anthocyanin pigments, inducers of Rhizobium nodulation genes, and antimicrobial phytoalexins. The 1.85 A resolution crystal structure of alfalfa CHI in complex with (2S)-naringenin reveals a novel open-faced beta-sandwich fold. Currently, proteins with homologous primary sequences are found only in higher plants. The topology of the active site cleft defines the stereochemistry of the cyclization reaction. The structure and mutational analysis suggest a mechanism in which shape complementarity of the binding cleft locks the substrate into a constrained conformation that allows the reaction to proceed with a second-order rate constant approaching the diffusion controlled limit. This structure raises questions about the evolutionary history of this structurally unique plant enzyme.
Organizational Affiliation:
Structural Biology Laboratory, The Salk Institute for Biological Studies, 10010 N. Torrey Pines Road, La Jolla, California 92037, USA.