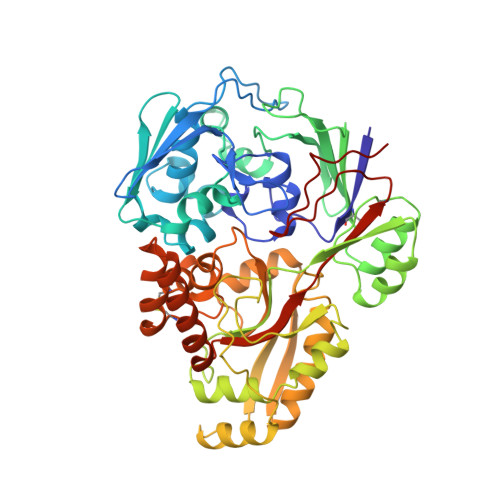
Crystal structure of the dipeptide binding protein from Escherichia coli involved in active transport and chemotaxis.
Dunten, P., Mowbray, S.L.(1995) Protein Sci 4: 2327-2334
- PubMed: 8563629
- DOI: https://doi.org/10.1002/pro.5560041110
- Primary Citation of Related Structures:
1DPP - PubMed Abstract:
The Escherichia coli periplasmic dipeptide binding protein functions in both peptide transport and taxis toward peptides. The structure of the dipeptide binding protein in complex with Gly-Leu (glycyl-L-leucine) has been determined at 3.2 A resolution. The binding site for dipeptides is designed to recognize the ligand's backbone while providing space to accommodate a variety of side chains. Some repositioning of protein side chains lining the binding site must occur when the dipeptide's second residue is larger than leucine. The protein's fold is very similar to that of the Salmonella typhimurium oligopeptide binding protein, and a comparison of the structures reveals the structural basis for the dipeptide binding protein's preference for shorter peptides.
Organizational Affiliation:
Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala, Sweden. dunten@xray.bmc.uu.se