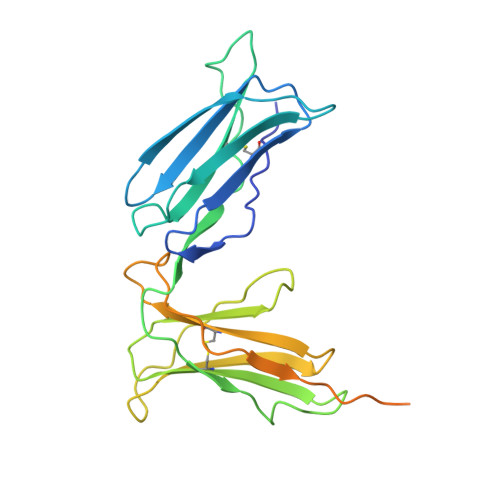
Crystal structure of the human p58 killer cell inhibitory receptor (KIR2DL3) specific for HLA-Cw3-related MHC class I.
Maenaka, K., Juji, T., Stuart, D.I., Jones, E.Y.(1999) Structure 7: 391-398
- PubMed: 10196125
- DOI: https://doi.org/10.1016/s0969-2126(99)80052-5
- Primary Citation of Related Structures:
1B6U - PubMed Abstract:
T cells and natural killer (NK) cells perform complementary roles in the cellular immune system. T cells identify infected cells directly through recognition of antigenic peptides that are displayed at the target cell surface by the classical major histocompatibility complex (MHC) class I molecules. NK cells monitor the target cell surface for malfunction of this display system, lysing potentially infected cells that might otherwise evade recognition by the T cells. Human killer cell inhibitory receptors (KIRs) control this process by either inhibiting or activating the cytotoxic activity of NK cells via specific binding to MHC class I molecules on the target cell.
Organizational Affiliation:
Laboratory of Molecular Biophysics, Rex Richards Building, South Parks Road, Oxford OX1 3QU, UK. katsumi@biop.ox.ac.uk